1.4.4. Advanced operations¶
Section contents
1.4.4.1. Polynomials¶
NumPy also contains polynomials in different bases:
For example, 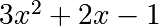 :
:
>>> p = np.poly1d([3, 2, -1])
>>> p(0)
-1
>>> p.roots
array([-1. , 0.33333333])
>>> p.order
2
>>> x = np.linspace(0, 1, 20)
>>> y = np.cos(x) + 0.3*np.random.rand(20)
>>> p = np.poly1d(np.polyfit(x, y, 3))
>>> t = np.linspace(0, 1, 200) # use a larger number of points for smoother plotting
>>> plt.plot(x, y, 'o', t, p(t), '-')
[<matplotlib.lines.Line2D object at ...>, <matplotlib.lines.Line2D object at ...>]
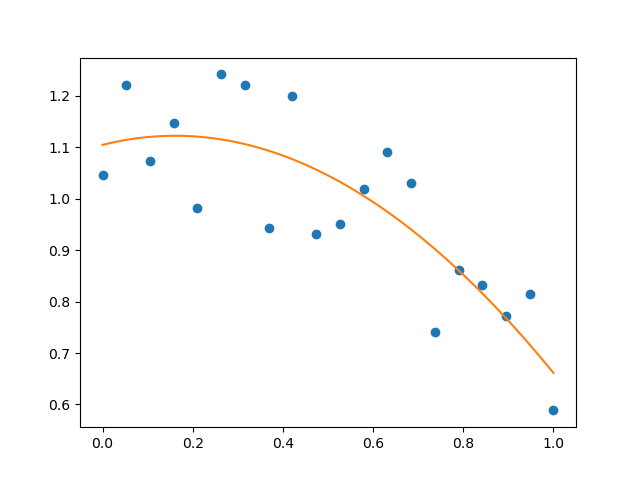
See http://numpy.org/doc/stable/reference/routines.polynomials.poly1d.html for more.
More polynomials (with more bases)¶
NumPy also has a more sophisticated polynomial interface, which supports e.g. the Chebyshev basis.
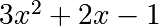 :
:
>>> p = np.polynomial.Polynomial([-1, 2, 3]) # coefs in different order!
>>> p(0)
-1.0
>>> p.roots()
array([-1. , 0.33333333])
>>> p.degree() # In general polynomials do not always expose 'order'
2
Example using polynomials in Chebyshev basis, for polynomials in
range [-1, 1]:
>>> x = np.linspace(-1, 1, 2000)
>>> y = np.cos(x) + 0.3*np.random.rand(2000)
>>> p = np.polynomial.Chebyshev.fit(x, y, 90)
>>> plt.plot(x, y, 'r.')
[<matplotlib.lines.Line2D object at ...>]
>>> plt.plot(x, p(x), 'k-', lw=3)
[<matplotlib.lines.Line2D object at ...>]
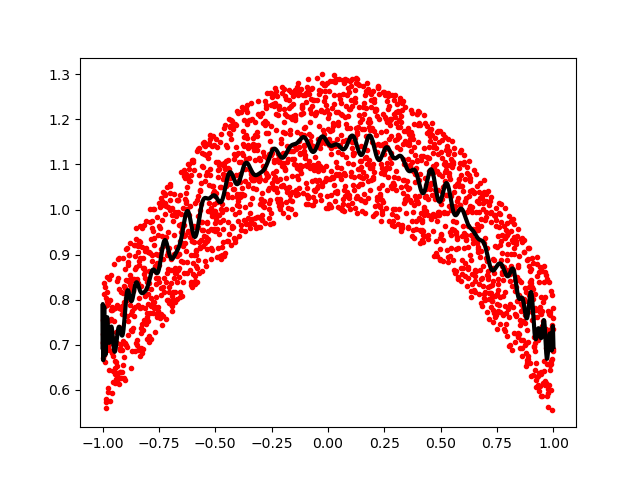
The Chebyshev polynomials have some advantages in interpolation.
1.4.4.2. Loading data files¶
Text files¶
Example: populations.txt:
# year hare lynx carrot 1900 30e3 4e3 48300 1901 47.2e3 6.1e3 48200 1902 70.2e3 9.8e3 41500 1903 77.4e3 35.2e3 38200
>>> data = np.loadtxt('data/populations.txt')
>>> data
array([[ 1900., 30000., 4000., 48300.],
[ 1901., 47200., 6100., 48200.],
[ 1902., 70200., 9800., 41500.],
...
>>> np.savetxt('pop2.txt', data)
>>> data2 = np.loadtxt('pop2.txt')
Note
If you have a complicated text file, what you can try are:
np.genfromtxt- Using Python’s I/O functions and e.g. regexps for parsing (Python is quite well suited for this)
Reminder: Navigating the filesystem with IPython
In [1]: pwd # show current directory
'/home/user/stuff/2011-numpy-tutorial'
In [2]: cd ex
'/home/user/stuff/2011-numpy-tutorial/ex'
In [3]: ls
populations.txt species.txt
Images¶
Using Matplotlib:
>>> img = plt.imread('data/elephant.png')
>>> img.shape, img.dtype
((200, 300, 3), dtype('float32'))
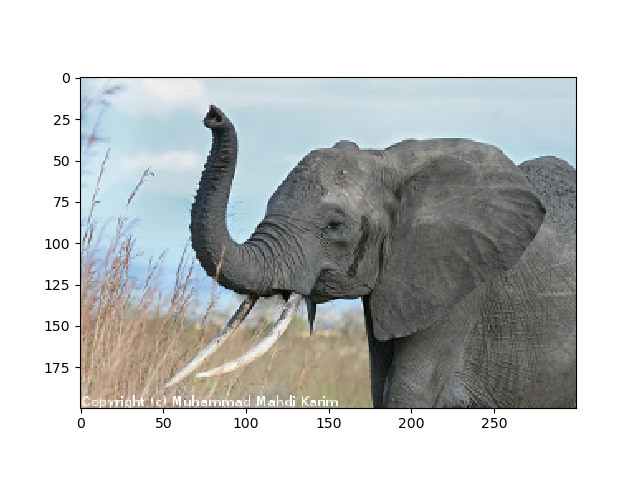
>>> plt.imshow(img)
<matplotlib.image.AxesImage object at ...>
>>> plt.savefig('plot.png')
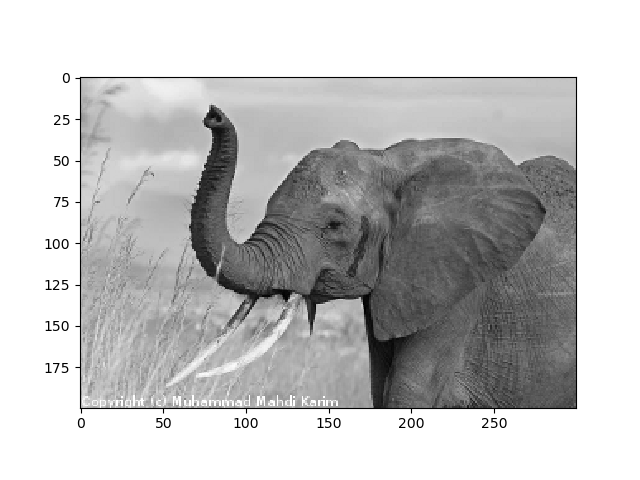
>>> plt.imsave('red_elephant.png', img[:,:,0], cmap=plt.cm.gray)
This saved only one channel (of RGB):
>>> plt.imshow(plt.imread('red_elephant.png'))
<matplotlib.image.AxesImage object at ...>
Other libraries:
>>> import imageio
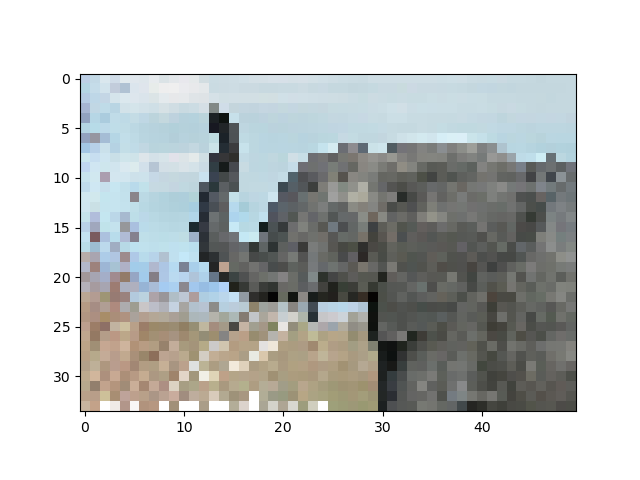
>>> imageio.imsave('tiny_elephant.png', img[::6,::6])
>>> plt.imshow(plt.imread('tiny_elephant.png'), interpolation='nearest')
<matplotlib.image.AxesImage object at ...>
NumPy’s own format¶
NumPy has its own binary format, not portable but with efficient I/O:
>>> data = np.ones((3, 3))
>>> np.save('pop.npy', data)
>>> data3 = np.load('pop.npy')
Well-known (& more obscure) file formats¶
- HDF5: h5py, PyTables
- NetCDF:
scipy.io.netcdf_file, netcdf4-python, … - Matlab:
scipy.io.loadmat,scipy.io.savemat - MatrixMarket:
scipy.io.mmread,scipy.io.mmwrite - IDL:
scipy.io.readsav
… if somebody uses it, there’s probably also a Python library for it.
Exercise: Text data files
Write a Python script that loads data from populations.txt:: and drop the last column and the first
5 rows. Save the smaller dataset to pop2.txt.
NumPy internals
If you are interested in the NumPy internals, there is a good discussion in Advanced NumPy.
